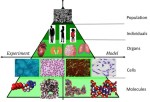
 During the lock-down in the UK due to the coronavirus pandemic, I have been reading about viruses and the modelling of them. It is a multi-disciplinary and multi-scale problem; so, something that engineers should be well-equipped to tackle. It is a multi-scale because we need to understand the spread of the virus in the human population so that we can control it, we need to understand the process of infection in individuals so that we can protect them, and we need to understand the mechanisms of virus-cell interaction so that we can stop the replication of the virus. At each size scale, models capable of representing the real-world processes will help us explore different approaches to arresting the progress of the virus and will need to be calibrated and validated against measurements. This can be represented in the sort of model-test pyramid shown in the top graphic that has been used in the aerospace industry [1-2] for many years [see ‘Hierarchical modelling in engineering and biology’ on March 14th, 2018] and which we have recently introduced in the nuclear fission [3] and fusion [4] industries [see ‘Thought leadership in fusion engineering’ on October 9th, 2019]. At the top of the pyramid, the spread of the virus in the population is being modelled by epidemiologists, such as Professor Neil Ferguson [5], using statistical models based on infection data. However, I am more interested in the bottom of the pyramid because the particles of the coronavirus are about the same size as the nanoparticles that I have been studying for some years [see ‘Slow moving nanoparticles’ on December 13th, 2017] and their motion appears to be dominated by diffusion processes [see ‘Salt increases nanoparticle diffusion’ on April 22nd, 2020] [6-7]. The first step towards virus infection of a cell is diffusion of the virus towards the cell which is believed to be a relatively slow process and hence a good model of diffusion would assist in designing drugs that could arrest or decelerate infection of cells [8]. Many types of virus on entering the cell make their way to the nucleus where they replicate causing the cell to die, afterwhich the virus progeny are dispersed to repeat the process. You can see part of this sequence for coronavirus (SARS-COV-2) in this sequence of images. The trafficking across the cytoplasm of the cell to the nucleus can occur in a number of ways including the formation of a capsule or endosome that moves across the cell towards the nuclear membrane where the virus particles leave the endosome and travel through microtubules into the nucleus. Holcman & Schuss [9] provide a good graphic illustrating these transport mechanisms. In 2019, Briane et al [10] reviewed models of diffusion of intracellular particles inside living eukaryotic cells, i.e. cells with a nuclear enclosed by a membrane as in all animals. Intracellular diffusion is believed to be driven by Brownian motion and by motor-proteins including dynein, kinesin and myosin that enable motion through microtubules. They observed that the density of the structure of cytoplasm, or cytoskeleton, can hinder the free displacement of a particle leading to subdiffusion; while, cytoskeleton elasticity and thermal bending can accelerate it leading to superdiffusion. These molecular and cellular interactions are happening at disparate spatial and temporal scales [11] which is one of the difficulties encountered in creating predictive simulations of virus-cell interactions. In other words, the bottom layers of the model-test pyramid appear to be constructed from many more strata when you start to look more closely. And, you need to add a time dimension to it. Prior to the coronavirus pandemic, more modelling efforts were perhaps focussed on understanding the process of infection by Human Immunodeficiency Virus (HIV), including by a multi-national group of scientists from Chile, France, Morocco, Russia and Spain [12-14]. However, the current coronavirus pandemic is galvanising researchers who are starting to think about novel ways of building multiscale models that encourage multidisciplinary collaboration by dispersed groups, [e.g. 15].
During the lock-down in the UK due to the coronavirus pandemic, I have been reading about viruses and the modelling of them. It is a multi-disciplinary and multi-scale problem; so, something that engineers should be well-equipped to tackle. It is a multi-scale because we need to understand the spread of the virus in the human population so that we can control it, we need to understand the process of infection in individuals so that we can protect them, and we need to understand the mechanisms of virus-cell interaction so that we can stop the replication of the virus. At each size scale, models capable of representing the real-world processes will help us explore different approaches to arresting the progress of the virus and will need to be calibrated and validated against measurements. This can be represented in the sort of model-test pyramid shown in the top graphic that has been used in the aerospace industry [1-2] for many years [see ‘Hierarchical modelling in engineering and biology’ on March 14th, 2018] and which we have recently introduced in the nuclear fission [3] and fusion [4] industries [see ‘Thought leadership in fusion engineering’ on October 9th, 2019]. At the top of the pyramid, the spread of the virus in the population is being modelled by epidemiologists, such as Professor Neil Ferguson [5], using statistical models based on infection data. However, I am more interested in the bottom of the pyramid because the particles of the coronavirus are about the same size as the nanoparticles that I have been studying for some years [see ‘Slow moving nanoparticles’ on December 13th, 2017] and their motion appears to be dominated by diffusion processes [see ‘Salt increases nanoparticle diffusion’ on April 22nd, 2020] [6-7]. The first step towards virus infection of a cell is diffusion of the virus towards the cell which is believed to be a relatively slow process and hence a good model of diffusion would assist in designing drugs that could arrest or decelerate infection of cells [8]. Many types of virus on entering the cell make their way to the nucleus where they replicate causing the cell to die, afterwhich the virus progeny are dispersed to repeat the process. You can see part of this sequence for coronavirus (SARS-COV-2) in this sequence of images. The trafficking across the cytoplasm of the cell to the nucleus can occur in a number of ways including the formation of a capsule or endosome that moves across the cell towards the nuclear membrane where the virus particles leave the endosome and travel through microtubules into the nucleus. Holcman & Schuss [9] provide a good graphic illustrating these transport mechanisms. In 2019, Briane et al [10] reviewed models of diffusion of intracellular particles inside living eukaryotic cells, i.e. cells with a nuclear enclosed by a membrane as in all animals. Intracellular diffusion is believed to be driven by Brownian motion and by motor-proteins including dynein, kinesin and myosin that enable motion through microtubules. They observed that the density of the structure of cytoplasm, or cytoskeleton, can hinder the free displacement of a particle leading to subdiffusion; while, cytoskeleton elasticity and thermal bending can accelerate it leading to superdiffusion. These molecular and cellular interactions are happening at disparate spatial and temporal scales [11] which is one of the difficulties encountered in creating predictive simulations of virus-cell interactions. In other words, the bottom layers of the model-test pyramid appear to be constructed from many more strata when you start to look more closely. And, you need to add a time dimension to it. Prior to the coronavirus pandemic, more modelling efforts were perhaps focussed on understanding the process of infection by Human Immunodeficiency Virus (HIV), including by a multi-national group of scientists from Chile, France, Morocco, Russia and Spain [12-14]. However, the current coronavirus pandemic is galvanising researchers who are starting to think about novel ways of building multiscale models that encourage multidisciplinary collaboration by dispersed groups, [e.g. 15].
References
[1] Harris GL, Computer models, laboratory simulators, and test ranges: meeting the challenge of estimating tactical force effectiveness in the 1980’s, US Army Command and General Staff College, May 1979.
[2] Trevisani DA & Sisti AF, Air Force hierarchy of models: a look inside the great pyramid, Proc. SPIE 4026, Enabling Technology for Simulation Science IV, 23 June 2000.
[3] Patterson EA, Taylor RJ & Bankhead M, A framework for an integrated nuclear digital environment, Progress in Nuclear Energy, 87:97-103, 2016.
[4] Patterson EA, Purdie S, Taylor RJ & Waldon C, An integrated digital framework for the design, build and operation of fusion power plants, Royal Society Open Science, 6(10):181847, 2019.
[5] Verity R, Okell LC, Dorigatti I, Winskill P, Whittaker C, Imai N, Cuomo-Dannenburg G, Thompson H, Walker PGT, Fu H, Dighe A, Griffin JT, Baguelin M, Bhatia S, Boonyasiri A, Cori A, Cucunubá Z, FitzJohn R, Gaythorpe K, Green W, Hamlet A, Hinsley W, Laydon D, Nedjati-Gilani G, Riley S, van Elsland S, Volz E, Wang H, Wang Y, Xi X, Donnelly CA, Ghani AC, Ferguson NM, Estimates of the severity of coronavirus disease 2019: a model-based analysis., Lancet Infectious Diseases, 2020.
[6] Coglitore D, Edwardson SP, Macko P, Patterson EA, Whelan MP, Transition from fractional to classical Stokes-Einstein behaviour in simple fluids, Royal Society Open Science, 4:170507, 2017.
[7] Giorgi F, Coglitore D, Curran JM, Gilliland D, Macko P, Whelan M, Worth A & Patterson EA, The influence of inter-particle forces on diffusion at the nanoscale, Scientific Reports, 9:12689, 2019.
[8] Gilbert P-A, Kamen A, Bernier A & Garner A, A simple macroscopic model for the diffusion and adsorption kinetics of r-Adenovirus, Biotechnology & Bioengineering, 98(1):239-251,2007.
[9] Holcman D & Schuss Z, Modeling the early steps of viral infection in cells, Chapter 9 in Stochastic Narrow Escape in Molecular and Cellular Biology, New York: Springer Science+Business Media, 2015.
[10] Braine V, Vimond M & Kervrann C, An overview of diffusion models for intracellular dynamics analysis, Briefings in Bioinformatics, Oxford University Press, pp.1-15, 2019.
[11] Holcman D & Schuss Z, Time scale of diffusion in molecular and cellular biology, J. Physics A: Mathematical and Theoretical, 47:173001, 2014.
[12] Bocharov G, Chereshnev V, Gainov I, Bazhun S, Bachmetyev B, Argilaguet J, Martinez J & Meyerhans A, Human immunodeficiency virus infection: from biological observations to mechanistic mathematical modelling, Math. Model. Nat. Phenom., 7(5):78-104, 2012.
[13] Bocharov G, Meyerhans A, Bessonov N, Trofimchuk S & Volpert V, Spatiotemporal dynamics of virus infection spreading in tissues, PLOS One, 11(12):e)168576, 2016.
[14] Bouchnita A, Bocharov G, Meyerhans A & Volpert V, Towards a multiscale model of acute HIV infection, Computation, 5(6):5010006, 2017.
[15] Sego TJ, Aponte-Serrano JO, Ferrari-Gianlupi J, Heaps S, Quardokus EM & Glazier JA, A modular framework for multiscale spatial modeling of viral infection and immune respons in epithelial tissue, bioRxiv. 2020.
 In 2021 Emily Bender and her colleagues published a paper suggesting that the Large Language Models (LLMs) underpinning many Artificial Intelligence applications (AI apps) were little more than stochastic parrots. They described LLMs as ‘a system for haphazardly stitching together sequences of linguistic forms it has observed in its vast training data, according to probabilistic information about how they combine, but without any reference to meaning’. This has fuelled the ongoing debate about the real capabilities of AI apps versus the hype from the companies trying persuade us to use them. Most AI apps are based on statistical analysis of data as stated by Bender et al; however, there is a trend toward physics-based machine learning in which known laws of physics are combined with machine-learning algorithms trained on data sets [see for example the recent review by Meng et al, 2025]. We have been fitting data to models for a very long time. In the fifth century BC, the Babylonians made perhaps one of the greatest breakthroughs in the history of science, when they realized that mathematical models of astronomical motion could be used to extrapolate data and make predictions. They had been recording astronomical observations since 3400 BC and the data was all collated in cuneiform in the library at Nineveh belonging to King Ashurbanipal who ruled from 669-631 BC. While our modern-day digital storage capacity in data centres might far exceed the clay tablets with cuneiform symbols found in Ashurbanipal’s library, it seems unlikely that they will survive five thousand years as part of the data from the Babylonians’ astronomical observations has done and still be readable.
In 2021 Emily Bender and her colleagues published a paper suggesting that the Large Language Models (LLMs) underpinning many Artificial Intelligence applications (AI apps) were little more than stochastic parrots. They described LLMs as ‘a system for haphazardly stitching together sequences of linguistic forms it has observed in its vast training data, according to probabilistic information about how they combine, but without any reference to meaning’. This has fuelled the ongoing debate about the real capabilities of AI apps versus the hype from the companies trying persuade us to use them. Most AI apps are based on statistical analysis of data as stated by Bender et al; however, there is a trend toward physics-based machine learning in which known laws of physics are combined with machine-learning algorithms trained on data sets [see for example the recent review by Meng et al, 2025]. We have been fitting data to models for a very long time. In the fifth century BC, the Babylonians made perhaps one of the greatest breakthroughs in the history of science, when they realized that mathematical models of astronomical motion could be used to extrapolate data and make predictions. They had been recording astronomical observations since 3400 BC and the data was all collated in cuneiform in the library at Nineveh belonging to King Ashurbanipal who ruled from 669-631 BC. While our modern-day digital storage capacity in data centres might far exceed the clay tablets with cuneiform symbols found in Ashurbanipal’s library, it seems unlikely that they will survive five thousand years as part of the data from the Babylonians’ astronomical observations has done and still be readable.


